Role of epigenomics and gene expression in the regulation of immune responses to WPBR in sugar pine

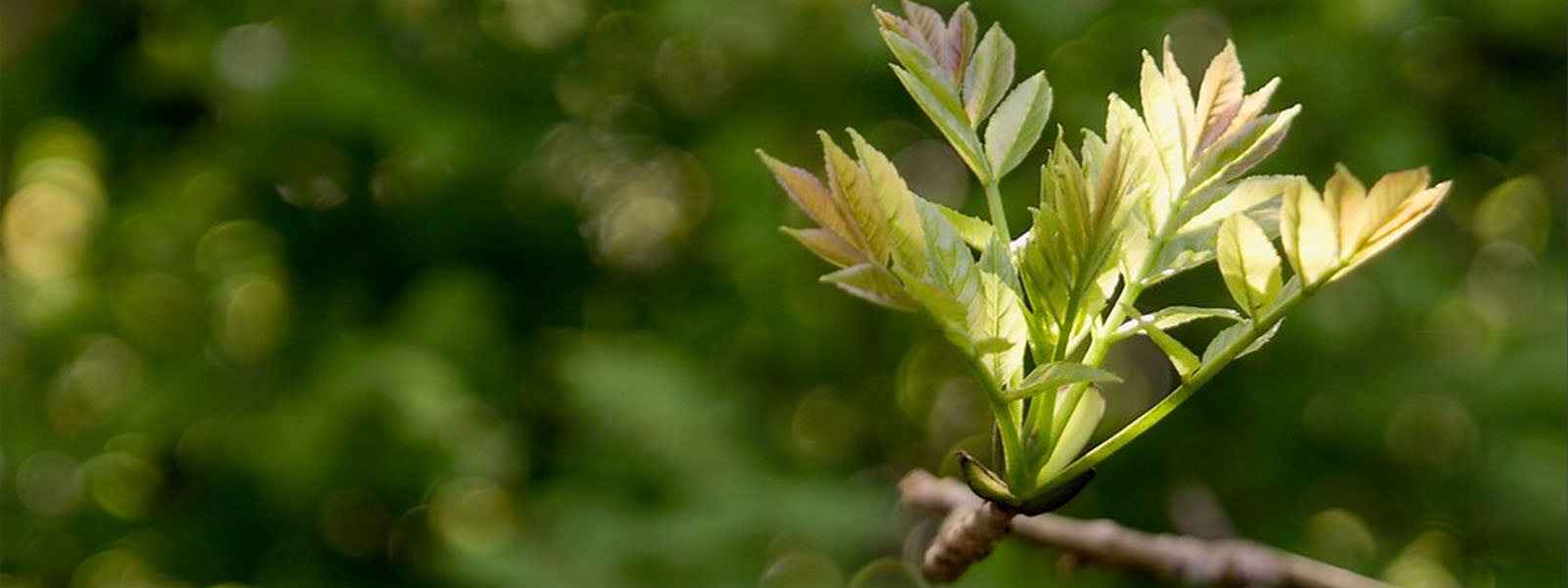
In highly-repetitive, densely-methylated conifer genomes, DNA methylation may play an important role in the response to pathogens, as varying levels of resistance and high levels of phenotypic plasticity are observed in natural populations of conifer species. Conifers possess some distinctive features in the RdDM pathway, such as low frequency of 24nt sRNAs, low numbers of Dicer-like 3 (DCL3) proteins, MIR loci composed by many long-terminal repeat-retrotransposons (LTR-RTs), and a high frequency of 21nt sRNAs. However, huge and complex draft genomes, limited genome-wide studies, outcrossing populations, and long-generation times have complicated the study of transgenerational inheritance. This project will investigate the White Pine Blister Rust (WPBR) - sugar pine pathosystem to understand the role of DNA methylation in regulating trans-generational immune responses. White pine blister rust (WPBR) is a devastating fungal disease causing great economic and ecological loss in five-needle pines such as sugar pine (Pinus lambertiana) in North America. In our previous studies in the pathosystem, we genotyped a large number of individuals for the presence or absence of resistance (major gene resistance, MGR and quantitative). These individuals were cloned and grown in contrasting environments for the last 20-30 years. To further examine resistance in sugar pine, we will construct a high-resolution map of genome-wide DNA methylation and examine whole-genome DNA methylation and expression in the progeny of resistant and susceptible trees after initial pathogen infection. The complex and fragmented genome assembly will be enhanced through long-read technologies and Hi-C. The resulting improvements in contiguity and accuracy will be paired with transcriptomic technologies, PacBio Iso-Seq. This will result in an enhanced understanding of transgenerational epigenetics, which could be used to reduce the very long time (~20 years) required by traditional breeding to generate resistant individuals. In specific, we aim to 1) Assess whether the RdDM pathway activity is reduced during maturity due to a decrease in the frequency of 24nt sRNAs, resulting in increased TE activity; 2) Identify the parental’ environmental factors (biotic and abiotic) that lead to hypomethylation resulting in heritable increased expression of NBS-LRR resistance genes in the offspring; 3) Evaluate the changes in DNA methylation and expression of NBS-LRR and other MGR resistance-related genes during different infection stages.
Team:
Susan McEvoy,
Akriti Bhattarai,
Emily Lesinski
Collaboration with:
Amanda De La Torre (NAU),
Richard Sniezko (USDA),
Laura Figueroa Corona (NAU),
Aleksey Zimin (Johns Hopkins)
Investigating Genetic Variation in Green Ash to Reduce Tree Mortality Against EAB

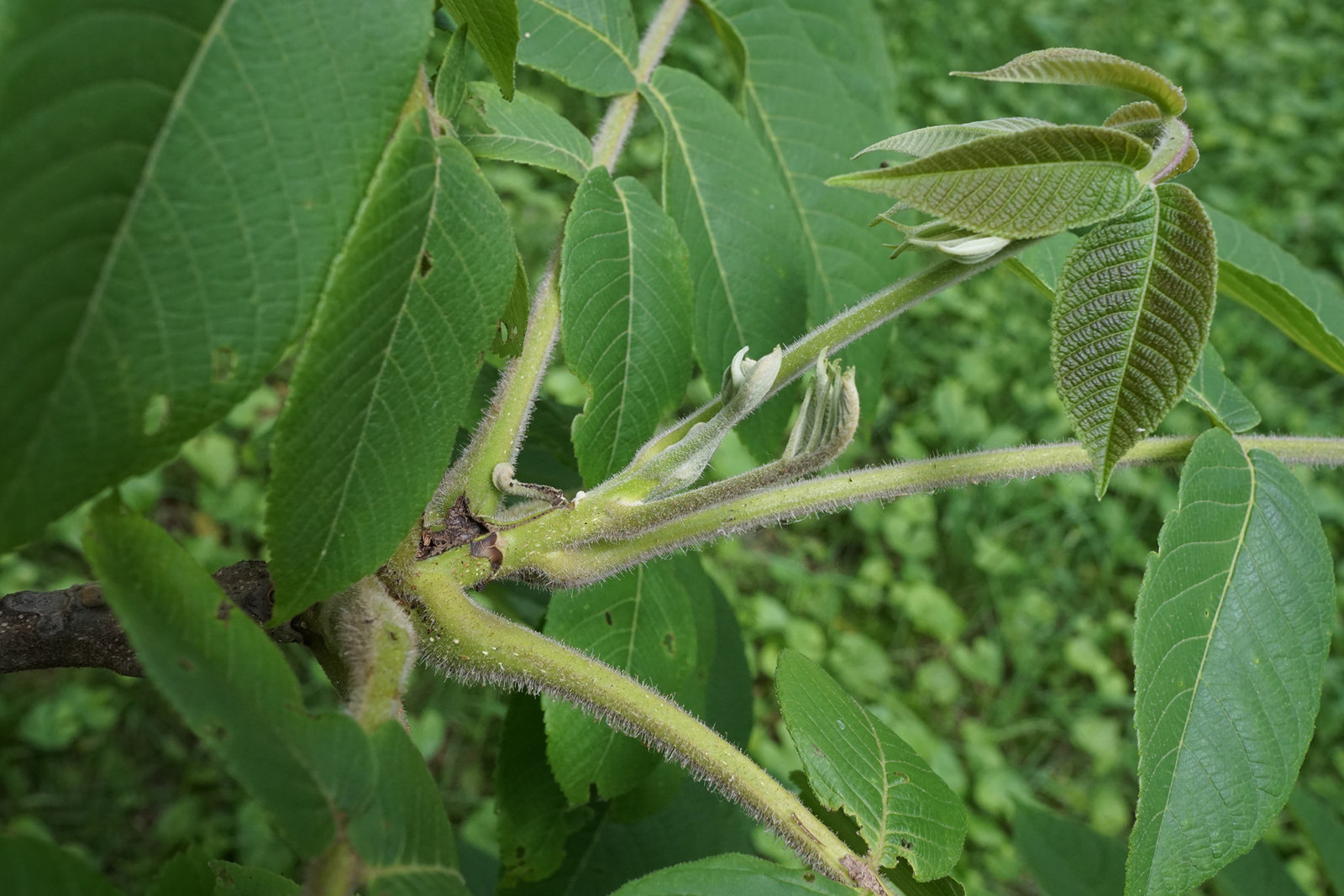
With the recent invasion of emerald ash borer (EAB) in North America, the health and quantity of American ash (genus: Fraxinus) trees has never been more at risk. Targeting every member of the Fraxinus genus, including Fraxinus americana (white ash), Fraxinus pennsylvanica (green ash), and Fraxinus nigra (black ash), EAB is proving to be detrimental to the American ash population as a whole. It was introduced to the United States from Asia through international trade, and now the invasive pest targets both healthy and unhealthy Fraxinus ash trees, feeds on their nutrient channels, and ultimately causes death in less than 3 years with a near 100% kill rate. In just five years after its initial discovery in 2002, EAB had destroyed more than 53 million native ash trees, and has been rapidly spreading throughout New England and as far west as Colorado. With the vast economic and ecological impacts the total destruction of this natural resource could have, researchers are under immense pressure to discover methods of protecting these species. We will study the impact of EAB through informed sampling of impacted populations and perform a genome-wide association study (ddRADSeq). The variants will be investigated with phenotypes assessed by our colleagues (metabolomics, growth traits, and disease resistance) While it is unlikely we can identify full resistance to this high mortality pest, we hope to uncover potential genes in lingering ash that may reduce susceptibility.
Team:
Jeremy Bennett,
Ava Fritz,
Irene Cobo Simon
Collaboration with:
Jennifer Koch (USDA),
Jeanne Romero-Severson (Notre Dame U.)
Comparative genomics in hardwoods: assembly and annotation of three maples

Sugar maple, a long-lived deciduous hardwood native to northeastern North America, is a dominant species in temperate forests. This shade-tolerant tree is present at low to mid elevations and has a continuous distribution from southern Quebec to the southeastern United States. Sugar maples promote Nitrogen mineralization, reduce nitrate load in groundwater, and generate beneficial nutrients for the soil. They are also an iconic species associated with vibrant Autumn hues. From an economic perspective, sugar maple may be the most valuable member of northeastern temperate forests as the primary source of maple syrup as well as hardwood timber. During the past decades, sugar maple decline increased across the natural range, with notable reductions in northeastern forests. Declining populations are characterized by a loss of crown vigor, dieback of fine branches, reduced radial growth, and associated low regeneration. Inconsistent regional patterns in both managed and unmanaged forests made the source of sugar maple decline elusive despite substantial research. Adaptive genetic variation in relevant genes and phenotypic plasticity are essential for survival in future conditions. Predominantly outcrossing, sugar maple populations are long lived and sessile, making them ideal for associating genetic diversity with environmental metrics. Despite this advantage, genomic resources for hardwood species have been slow to develop. Their evolutionary history contributes to divergent chromosomal architectures and multiple whole-genome duplications and rearrangements. These qualities introduce complexities in the assembly and annotation of reference sequences. Reduced cost and increased availability of long read technologies has allowed for the recent efforts to sequence, assemble, and annotate three Acer species to conduct comparative genomics analysis and further understand their adaptive potential.
Team:
Susan McEvoy
Collaboration with: Nathan Swenson (UMD),
Alex Touern-Trend (UConn),
Uzay Sezen (UConn),
Paul Schaberg (USDA)
Not so evergreen: investigating leaf senescence in a deciduous conifer
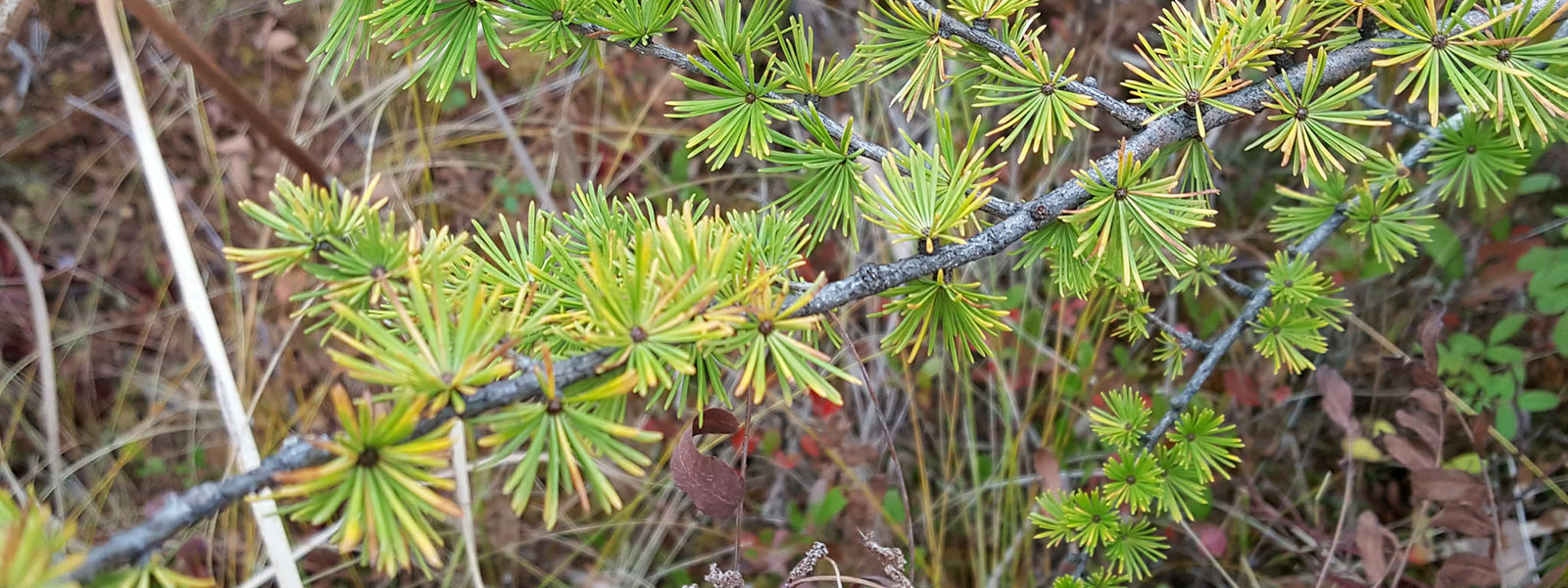
 Eastern larch (or tamarack) is one species among a handful of deciduous conifers. There are only 19 gymnosperms that undergo Autumn leaf senescence, and most belong to the Larix (larch) genus. While the majority of conifers shed their needles at varying intervals over their lifespan (4-5 years), tamaracks behave like many deciduous angiosperms, displaying a deep yellow hue in early to mid Autumn and fully losing all of their needles by winter. Leaf (or needle) senescence is considered the final stage in leaf development, and is associated with cellular death. This process is highly coordinated as there are parallel changes in metabolism and cell structure. A better understanding of Autumn leaf senescence in gymnosperms can provide further insight on their evolutionary history. The abscission zone (tissue providing the attachment of the needle to the stem) was sampled and sequenced for a replicated timecourse study to identify gene expression changes through seasonal senescence. We are currently investigating the pathways and genes that are well conserved with broad leaf angiosperms and those that are unique to conifers.
Lead: Kristiana Stover
Contributors: Vidya Vuruputoor
Eastern larch (or tamarack) is one species among a handful of deciduous conifers. There are only 19 gymnosperms that undergo Autumn leaf senescence, and most belong to the Larix (larch) genus. While the majority of conifers shed their needles at varying intervals over their lifespan (4-5 years), tamaracks behave like many deciduous angiosperms, displaying a deep yellow hue in early to mid Autumn and fully losing all of their needles by winter. Leaf (or needle) senescence is considered the final stage in leaf development, and is associated with cellular death. This process is highly coordinated as there are parallel changes in metabolism and cell structure. A better understanding of Autumn leaf senescence in gymnosperms can provide further insight on their evolutionary history. The abscission zone (tissue providing the attachment of the needle to the stem) was sampled and sequenced for a replicated timecourse study to identify gene expression changes through seasonal senescence. We are currently investigating the pathways and genes that are well conserved with broad leaf angiosperms and those that are unique to conifers.
Lead: Kristiana Stover
Contributors: Vidya Vuruputoor,
Irene Cobo-Simón,
Alison Scott,
Cynthia Webster,
M. Amee,
Olivia Maher
Associative transcriptomics and metagenomics to evaluate adaptation to acid rain in two hardwood species
 Understanding the population genetic structure, and gene expression patterns as it relates to different soil conditions can predict future trajectories of forest composition. No genetic studies have been carried out on the trees in the long-term ecological monitoring site, Hubbard Brook Experimental Forest (HBEF) in New Hampshire. Monitoring of growth performance in the field has revealed that sugar maple is on the decline. On the other hand, American beech is performing well in exacerbated cation depleted soils. Controlled field experiments have examined the effects of Ca and Al treatments when applied through the soil. Dominant sugar maple trees remained unaffected but non-dominant trees responded positively to Ca amendment. Transcriptomics of the plant tissues and metagenomics of the associated soil microbial/fungal communities are underway to build a more complete picture of forest response.
Lab Team: Alex Trouern-Trend
Collaboration with:
Uzay Sezen,
Paul Schaberg (US Forest Service),
Susan McEvoy (SLU)
Understanding the population genetic structure, and gene expression patterns as it relates to different soil conditions can predict future trajectories of forest composition. No genetic studies have been carried out on the trees in the long-term ecological monitoring site, Hubbard Brook Experimental Forest (HBEF) in New Hampshire. Monitoring of growth performance in the field has revealed that sugar maple is on the decline. On the other hand, American beech is performing well in exacerbated cation depleted soils. Controlled field experiments have examined the effects of Ca and Al treatments when applied through the soil. Dominant sugar maple trees remained unaffected but non-dominant trees responded positively to Ca amendment. Transcriptomics of the plant tissues and metagenomics of the associated soil microbial/fungal communities are underway to build a more complete picture of forest response.
Lab Team: Alex Trouern-Trend
Collaboration with:
Uzay Sezen,
Paul Schaberg (US Forest Service),
Susan McEvoy (SLU)
Eco-Evolutionary Dynamics Across Arctic Meta-Ecosystems (EVOME)
The Evolving MetaEcosystems (EvoME) Institute will study the effects of global climate change on Arctic ecosystems. Ecosystems are complex communities of species that have evolved with each other and their shared environment over long periods of time. Understanding how these systems change over time is a Grand Challenge in Biology that is made urgent and policy-relevant by rapid climate change. This is particularly true in the Arctic, which is warming at least three times faster than the global average. Arctic ecosystems are uniquely suited to their extreme environment, and they provide food and livelihoods for human communities. It is critical to know whether species and ecosystems can evolve to match the pace of change, or whether they might fall apart or muddle along in a reduced state. EvoME will bring together experts from across biological disciplines to generate new insights at every scale, from genes to landscapes. It will document natural responses of multiple species in rivers and streamside tundra environments and conduct large-scale experiments on the flow of energy and genes between ecosystems. EvoME will foster a new generation of biologists trained to think and work across disciplines, with special attention to increasing inclusion and retention of researchers from underrepresented backgrounds, by a cross-disciplinary and cross-institutional course, a research fund for students, and a Fellows program. Finally, it will bring journalists into the research process to create—and help researchers create—innovative media and stories through blogs, social media, and radio stories that bring EvoME’s integrated understanding to public audiences, including rural and Alaska Native communities.
Our team focuses on Salix (willow), Arctic grayling (Thymallus arcticus), and key aquatic invertebrates (mayflies, beetles) sampled along longitudinal gradients that span major shifts in climate, hydrology, and nutrient flow. Using whole-genome and reduced-representation sequencing, coupled with trait and environmental data, the project explores dispersal, gene flow, local adaptation, and eco-evolutionary feedbacks under climate change.
Lab Team: Brandon Lind,
Airianna McGuire,
Samira Obbu,
Mary Rutter
Collaborators: Linda Deegan (Woodwell Climate Research Center),
Mark Urban (UConn),
Natalie Boelman (Columbia University),
Jeff Lozier &
Carla Atkinson (University of Alabama),
Erik Schoen &
Peter Westley (University of Alaska Fairbanks) and
Rachel O’Neill ( University of Connecticut)
External websites:
https://www.woodwellclimate.org/project/evolving-meta-ecosystems-institute/
Pangenomics of Tsuga (Hemlock) to Uncover Mechanisms of Adelgid Resistance
This project develops a Tsuga pangenome to identify the genomic, structural, and regulatory variation associated with host defense to Hemlock Woolly Adelgid (HWA). By integrating long-read sequencing, structural variant detection, comparative genomics, and transcriptomics, we aim to pinpoint candidate resistance loci and evaluate their conservation across Tsuga species. This work supports breeding, conservation, and restoration strategies for this foundation forest genus under severe biotic threat.
Lab Team: Meghan Myles
Collaborators: Arnold Arboretum of Harvard University,
Karl Fetter (USDA)
Genomic Signatures of Extreme Longevity in Bristlecone Pines
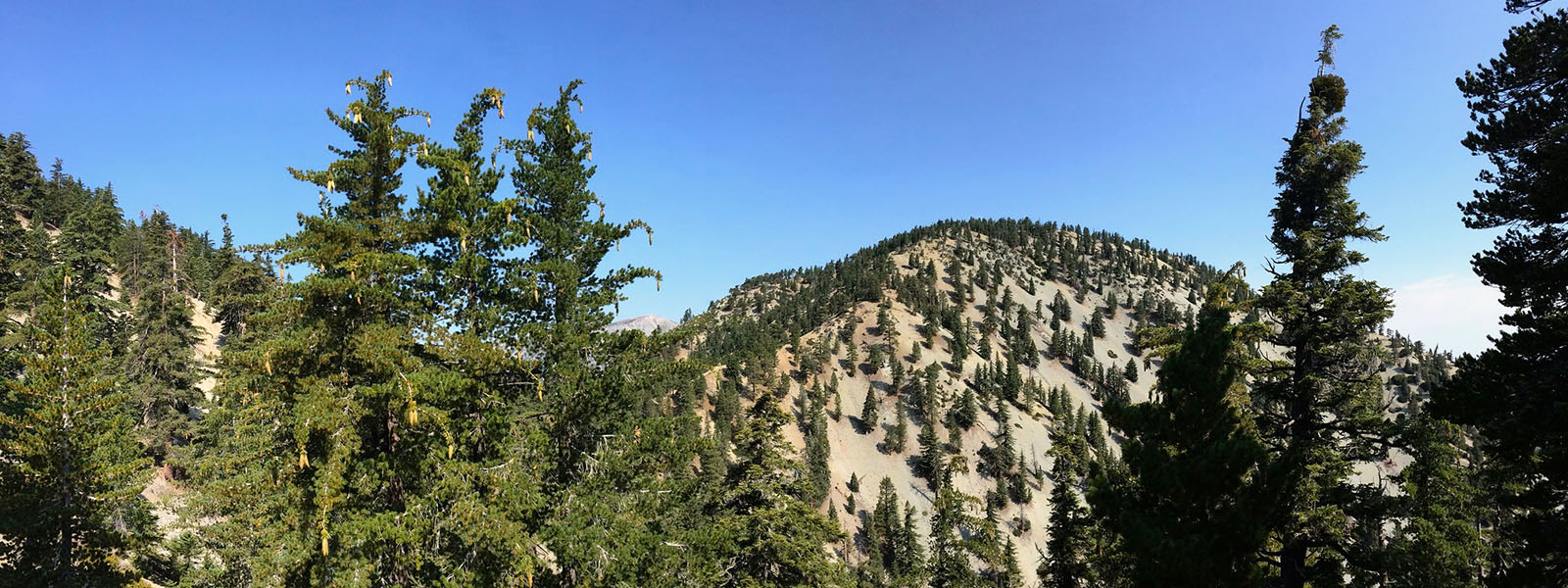
Bristlecone pines (Pinus longaeva) can live for more than 5,000 years, making them the longest-lived non-clonal organisms on Earth. This project leverages long-read genome sequencing, annotation, and comparative genomics across conifers to investigate the evolutionary basis of exceptional lifespan, cellular resilience, stress response, and maintenance pathways. Insights into genome stability, transposable element dynamics, and protective biochemical mechanisms aim to illuminate how perennial plants evade senescence over millennia.
Lab Team: Ruiwen Lin,
Meghan Myles
Collaborators: David Neale (UCDavis),
Aleksey Zimin (Johns Hopkins U.).
Steven Salzberg (Johns Hopkis U.), Winston Timp (Johns Hopkins U.)
Genomic Resource Development and Genotyping Tools for Fir (Abies) Species
This applied genomics initiative provides foundational resources for Abies, a genus of ecological and economic importance facing increasing climate- and pest-related pressures. We are assembling reference data, generating SNP-based genotyping tools, and building workflows to support association studies, breeding programs, and conservation planning. Outcomes will guide trait improvement, resilience assessment, and informed management decisions for fir species across North America.
Lab Team:
Vidya Vuruputoor,
Mary Rutter
Collaborators: Justin Whitehill (NCSU),
Ross Whetten (NCSU)
Epigenomic Variation and Adaptive Potential in White Pines
This work integrates methylome profiling, gene expression, and genomic variation to understand the role of epigenetic regulation in stress response and adaptation in white pines (Pinus spp.). Leveraging nanopore-based methylation data, the project explores how heritable and inducible epigenetic modifications shape local adaptation and may contribute to resilience in changing environments.
Lab Team: Emily Lesinkski
Collaborators: Amanda De La Torre (NAU),
Susan McEvoy,
Akriti Bhattarai
Genomic Resources and Resistance Mapping for Beech Bark Disease and Beech Leaf Disease
American beech (Fagus grandifolia) is threatened by two major diseases: Beech Bark Disease, a scale insect–fungal pathogen complex that has transformed eastern forests for over a century, and the emerging Beech Leaf Disease caused by Litylenchus crenatae ssp. mccannii. This project is developing comprehensive genomic resources including reference-quality datasets, transcriptomes and population resequencing to investigate host response, resistance mechanisms and adaptive potential. Fine-scale mapping and association studies are resolving a major QTL linked to BBD resistance, while complementary efforts are characterizing early defense signaling and resilience to BLD. These resources support marker-assisted resistance screening, inform deployment strategies and aim to safeguard the future of this keystone species through science-guided restoration.
Lab Team: Michelle Neitzey,
Airianna McGuire,
Cynthia Webster
Collaborators: Suzy Strickler (Chicago Botanic Garden),
Fei-Wei Lu (Boyce Thompson Institute), Susan McEvoy (SLU), Jennifer Koch (USDA), Dave Carey (USDA), David Burke (Holden Arboretum)
Leveraging omics to conserve Eastern hemlock, an imperiled conifer
Eastern hemlock (Tsuga canadensis) is a foundational tree species in eastern North American forests, yet it is rapidly declining due to infestation by the introduced hemlock woolly adelgid (HWA). With current biological and chemical control strategies offering only limited protection, there is a critical need to understand the genetic and physiological basis of natural tolerance. We generated the first high-quality, chromosome-scale reference genome for T. canadensis (16.28 Gb; 91% BUSCO completeness; 79% repetitive elements), establishing a foundational resource for conservation genomics and breeding programs. Leveraging this assembly, we investigated mechanisms of HWA resistance in the “Bulletproof” (BP) stand, a group of lingering hemlocks that persist despite long-term, high HWA pressure. To characterize defense responses, we integrated phenotypic, transcriptomic, and metabolomic data collected over one year from lingering and susceptible individuals across sites in Massachusetts, Pennsylvania, and North Carolina. Weighted gene co-expression network analyses (WGCNA) revealed modules consistently elevated in lingering trees, including pathways involved in terpenoid biosynthesis, cell wall remodeling, and immune and stress-response signaling. Broader pathway-level patterns highlighted differences in phenylpropanoid metabolism, seasonal stress-response regulation, and energy-related processes such as CoA and pantothenate pathways. Collectively, these results suggest that tolerance in lingering trees is maintained through dynamic, seasonally modulated defense strategies. This work provides new insight into the molecular architecture of HWA tolerance and delivers essential genomic tools to support restoration, resistance screening, and long-term conservation of eastern hemlock.
Team:
Vidya Vuruputoor,
Meghan Myles
Collaboration with:
Karl Fetter (USDA),
Tim Cernak (U, Michigan),
Jennifer Koch (USDA),
Rachel Kappler (Holden Arboretum)

The idea for CartograPlant began in June of 2011, when a group of forest tree biology researchers from different fields like physiology, ecology, genomics, and systematics realized the need for a unified platform to integrate and visualize spatial biological data. These researchers came together through workshops and collaborations funded by the NSF iPlant Collaborative (now CyVerse), a project designed to support data-driven biological research.
Their goal was to create a tool that could help bridge the gaps between disciplines and enable easier access to georeferenced population data, with traits and genotypes. The focus was on building a web-based application that could be deployed and used by researchers from various backgrounds, making complex data more accessible and useful for a wide range of studies.
The first version of CartograPlant (then, CartograTree) was released in 2012, built on the resources and infrastructure provided by iPlant. By 2015, a more refined version of the platform was launched, allowing users to identify, filter, compare, and visualize spatial data. The tool was designed to handle different types of datasets, including species distributions, genetic information, and environmental factors.
Today, CartograPlant continues to support the forest tree community, and increasingly, other plant systems as well. The application supports advanced analysis through HPC and reproducible workflows, such as population structure, genetic diversity, and association genetics. CarograPlant aims to help scientists understand how environmental and genetic factors influence the diversity and global distribution of plant species, engaging scientist and practitioners at all levels.
Software Access
Software Documentation
Lab Team:
Brandon Lind,
Meghan Myles,
Risharde Ramnath,
Emily Grau,
Gabe Barrett,
Vlad Savitsky,
Phoebe Zhou and
Trang Nguyen
Collaborators:
Meg Staton, Florence Caldwell, Chance Stribling (Department of Entomology and Plant Pathology, UTenn) and
Jonathan Rosenthal, Radka Wildova (Ecological Research Institute)

The TreeGenes Database traces its origins to the Dendrome Project, launched in the mid-1990s as one of the first USDA Agricultural Research Service (ARS) genome databases. Conceived as a centralized resource for forest tree genetics, Dendrome was developed to manage and share emerging molecular data for conifers and other tree species. It served as the first of three ARS genome databases, alongside those for maize and grasses, and provided early tools for the storage, retrieval, and visualization of genetic maps, expressed sequence tags (ESTs), and marker data.
Over time, Dendrome evolved into TreeGenes, expanding both its taxonomic scope and infrastructure to meet the demands of large-scale sequencing and comparative genomics. As high-throughput sequencing changed life science research, TreeGenes grew to accommodate large genetic datasets and diverse data types, from genome assemblies and transcriptomes to population genetics and environmental metadata.
A key development in TreeGenes’ evolution was its full adoption of the Tripal platform, an open-source toolkit that integrates the Chado schema with modern web content management (Drupal) to create flexible, interoperable biological databases. This transition enabled TreeGenes to modularize its infrastructure, enhance its scalability, and connect with other Tripal-based resources. Within this framework, TreeGenes develops and maintains custom Tripal modules, including CartograPlant, a map-based interface that visualizes genotypes, phenotypes, and environmental variables to support landscape and association genomics. The platform also integrates workflows that support metadata/data annotation, quality control, and analysis within a FAIR (Findable, Accessible, Interoperable, Reusable) data ecosystem.
Today, TreeGenes stands as one of the largest and longest running plant genomics databases in the world. It provides curated modules to access reference genome assemblies, transcriptomic data, variant calls, phenotypic observations, and environmental context, linked through standardized metadata and georeferenced accessions.
Software Access
Lab Team:
Brandon Lind,
Meghan Myles,
Gabe Barrett,
Risharde Ramnath,
Emily Grau,
Vlad Savitsky
Phoebe Zhou and
Trang Nguyen
Collaboration with:Meg Staton (Department of Entomology and Plant Pathology, UTenn)
Funding:
Publications:
Lind, B. M., Cobo-Simón, I., Myles, M., Barrett, G., Grau, E., Ramnath, R., Savitsky, V., Wegrzyn, J. L. (2025). CartograPlant: Bridging genomic, phenotypic, and environmental data to advance plant resilience and eco-evolutionary insight.
EcoEvoRxiv, 2025.09.30.
https://ecoevorxiv.org/repository/view/10352/
Wegrzyn, J. L., Falk, T., Grau, E., Buehler, S., Ramnath, R., & Herndon, N. (2019). Cyberinfrastructure and resources to enable an integrative approach to studying forest trees. Evolutionary Applications, 13(1), 228–241.
https://doi.org/10.1111/eva.12860
Spoor, S., Cheng, C. H., Sanderson, L. A., Condon, B., Almsaeed, A., Chen, M., Bretaudeau, A., Rasche, H., Jung, S., Main, D., Bett, K., Staton, M., Wegrzyn, J. L., Feltus, F. A., and Ficklin, S. P. (2019). Tripal v3: an ontology-based toolkit for construction of FAIR biological community databases.
Database, baz077.
https://doi.org/10.1093/database/baz077
Wegrzyn, J. L., Staton, M. A., Street, N. R., Main, D., Grau, E., Herndon, N., Buehler, S., Falk, T., Zaman, S., Ramnath, R., Richter, P., Sun, L., Condon, B., Almsaeed, A., Chen, M., Mannapperuma, C., Jung, S., and Ficklin, S. (2019). Cyberinfrastructure to improve forest health and productivity: The role of tree databases in connecting genomes, phenomes, and the environment.
Frontiers in Plant Science, 10:813.
https://doi.org/10.3389/fpls.2019.00813
Falk, T., Herndon, N., Grau, E., Buehler, S., Richter, P., Zaman, S., Baker, E. M., Ramnath, R., Ficklin, S., Staton, M., Feltus, F. A., Jung, S., Main, D., and Wegrzyn, J. L. (2018). Growing and cultivating the forest genomics database, TreeGenes.
Database, 2018: bay084-bay084. Retrieved from
http://dx.doi.org/10.1093/database/bay084
Harper, L., Campbell, J., Cannon, E. K. S., Jung, S., Poelchau, M., Walls, R., Andorf, C., Arnaud, E., Berardini, T. Z., Birkett, C., Cannon, S., Carson, J., Condon, B., Cooper, L., Dunn, N., Elsik, C. G., Farmer, A., Ficklin, S. P., Grant, D., Grau, E., Herndon, N., Hu, Z., Humann, J., Jaiswal, P., Jonquet, C., Laporte, M., Larmande, P., Lazo, G., McCarthy, F., Menda, N., Mungall, C., Munoz-Torres, M. C., Naithani, S., Nelson, R., Nesdill, D., Park, C., Reecy, J., Reiser, L., Sanderson, L., Sen, T. Z., Staton, M., Subramaniam, S., Tello-Ruiz, M. K., Unda, V., Unni, D., Wang, L., Ware, D., Wegrzyn, J. L., Williams, J., Woodhouse, M., Yu, J., and Main, D. (2018). AgBioData consortium recommendations for sustainable genomics and genetics databases for agriculture.
Database, bay088.
http://dx.doi.org/10.1093/database/bay088
Wegrzyn, J. L., Main, D., Figueroa, B., Choi, M., Yu, J., Neale, D. B., Stanton, M., Zheng, P., Ficklin, S., Cho, I., Peace, C., Evans, K., Volk, G., Oraguzie, N., Chen, C., Jr., F. G., Abbott, A. G. (2012). Uniform standards for genome databases in forest and fruit trees.
Tree Genetics and Genomes, 8(3): 549–557.
https://doi.org/10.1007/s11295-012-0494-7
 EnTAP is designed to improve the accuracy, speed, and flexibility of functional gene annotation. EnTAP integrates taxonomic scope to optimize the selection of the most appropriate descriptors, as well as filter for contaminants common in transcriptomes and genomes. EnTAP addresses the challenges associated with de novo transcriptome assembly that lead to inflated and inaccurate transcripts through target transcript coverage (RSEM expression estimates) and the prediction of viable open reading frames (TransDecoder). Following filters applied through assessment of true expression and frame selection, translated proteins are compared to up to three protein databases, and independently assigned to gene families via EggNOG. Sequence similarity is implemented through Diamond for rapid assessment, and Gene Ontology terms are assigned from a combination of high quality UniProt alignments, when available, and EggNOG. EnTAP can process header information from both EBI and NCBI databases to aid in the selection of a single alignment that results from a combination of weighted metrics describing similarity search score, taxonomic relationship, and informativeness.
Software Access
Software Documentation
Lab Team:
EnTAP is designed to improve the accuracy, speed, and flexibility of functional gene annotation. EnTAP integrates taxonomic scope to optimize the selection of the most appropriate descriptors, as well as filter for contaminants common in transcriptomes and genomes. EnTAP addresses the challenges associated with de novo transcriptome assembly that lead to inflated and inaccurate transcripts through target transcript coverage (RSEM expression estimates) and the prediction of viable open reading frames (TransDecoder). Following filters applied through assessment of true expression and frame selection, translated proteins are compared to up to three protein databases, and independently assigned to gene families via EggNOG. Sequence similarity is implemented through Diamond for rapid assessment, and Gene Ontology terms are assigned from a combination of high quality UniProt alignments, when available, and EggNOG. EnTAP can process header information from both EBI and NCBI databases to aid in the selection of a single alignment that results from a combination of weighted metrics describing similarity search score, taxonomic relationship, and informativeness.
Software Access
Software Documentation
Lab Team: Alex Hart,
Cynthia Webster and
Vidya Vuruputoor

U.S National Science Foundation
<b